Project Overview
This project develops artificial intelligence methods for automated single-cell annotation and spatial transcriptomics analysis. By leveraging advanced deep learning techniques, we aim to accelerate cellular phenotype identification, uncover spatial organization patterns in tissues, and provide computational tools that enable researchers to extract deeper biological insights from high-dimensional genomic data.
Goals
- Automated Cell Type Annotation: Develop AI models for accurate and rapid cell type identification in single-cell RNA sequencing data
- Spatial Pattern Discovery: Create algorithms to identify and characterize spatial cellular organization and cell-cell interactions
- Multi-Modal Integration: Build frameworks that integrate single-cell transcriptomics with spatial information for comprehensive tissue analysis
- Therapeutic Target Identification: Apply AI to discover cellular states and spatial niches relevant to disease progression and treatment resistance
Key Research Areas
Single-Cell RNA Sequencing Analysis
- Deep Learning for Cell Annotation: Transformer-based and graph neural network architectures for automated cell type classification
- Cell State Identification: Detection of rare cell populations, transitional states, and therapy-resistant subpopulations
- Transfer Learning: Leveraging large reference datasets to enable accurate annotation across tissues and disease contexts
- Trajectory Inference: AI-guided reconstruction of cellular differentiation and state transition pathways
Spatial Transcriptomics
- Spatial Pattern Recognition: Identifying tissue architecture, cellular neighborhoods, and microenvironmental niches
- Cell-Cell Interaction Modeling: Predicting ligand-receptor interactions and intercellular communication networks
- Spatial Domain Segmentation: Unsupervised and semi-supervised methods for tissue region identification
- Multi-Scale Analysis: Integration of cellular and tissue-level spatial features for comprehensive understanding
Applications in Cancer Biology
Our methods are particularly focused on understanding cancer heterogeneity and therapeutic resistance:
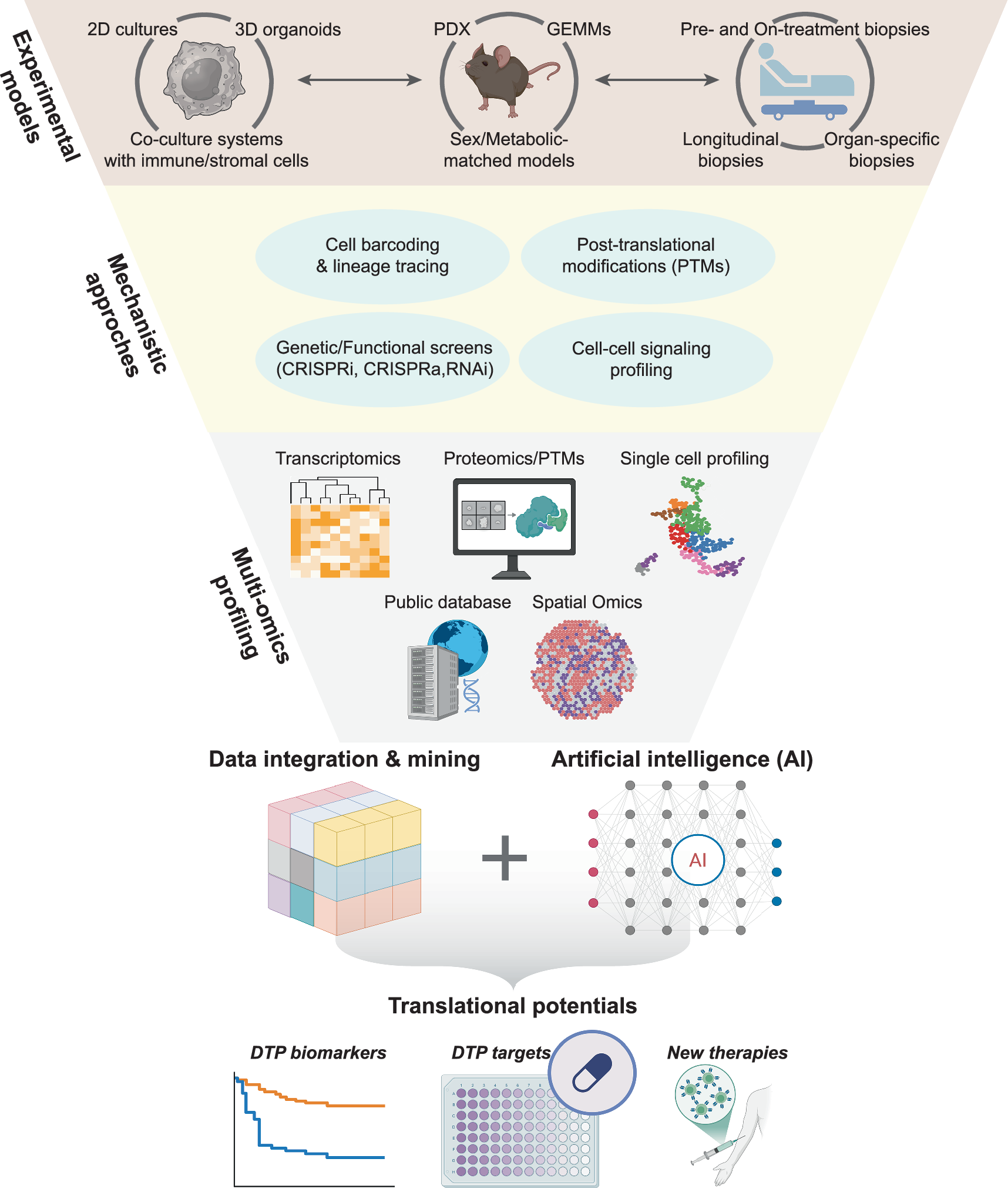
- Drug-Tolerant Persister Cells: Identifying rare persister cell populations that survive therapy and drive relapse
- Tumor Microenvironment Mapping: Characterizing immune, stromal, and malignant cell interactions in spatial context
- Treatment Response Prediction: Using single-cell and spatial features to predict therapeutic outcomes
- Biomarker Discovery: Discovering cellular signatures and spatial patterns associated with disease progression
Image reference: Adapted from Wang, Z., Wang, M., Dong, B. et al. “Drug-tolerant persister cells in cancer: bridging the gaps between bench and bedside.” Nature Communications 16, 10048 (2025). https://doi.org/10.1038/s41467-025-66376-6
Methodology
- Deep Neural Networks: Attention mechanisms, graph neural networks, and variational autoencoders for cellular representation learning
- Foundation Models: Adapting large-scale pre-trained models for single-cell and spatial transcriptomics
- Explainable AI: Interpretable architectures that reveal biological mechanisms underlying predictions
- Benchmark Development: Creating standardized datasets and evaluation metrics for method comparison
Team Members
Principal Investigators
- Dr. Chien Truong-Quoc - Hanoi University of Science and Technology (HUST)
- Dr. Nguyen Phuong Thuy - Hanoi Medical University
- Dr. Nguyen Minh Quan - Seoul National University
Students
- Graduate and undergraduate students specializing in computational biology, bioinformatics, and machine learning
Future Directions
- Multi-Omics Integration: Combining transcriptomics with proteomics, epigenomics, and metabolomics data
- Real-Time Analysis Platforms: Developing user-friendly web tools for rapid single-cell and spatial data analysis
- Clinical Translation: Applying computational methods to patient samples for precision medicine applications
- Cross-Species Analysis: Extending frameworks to enable comparative studies across model organisms and human tissues
Publications and Collaborations
Research findings are published in leading bioinformatics and computational biology journals. We actively collaborate with experimental biologists and clinicians to validate and apply our computational methods to real-world biological and medical questions.
Contact
For collaboration opportunities or research inquiries, please contact the AiRA Laboratory at Hanoi University of Science and Technology.
🔍 Search for AI for Single-Cell and Spatial Transcriptomics related papers on the Research page